Overview and implementation of clustering algorithm using the k-means technique.
%matplotlib inline
import matplotlib
import matplotlib.pyplot as plt
import numpy as np
from clustering__utils import *
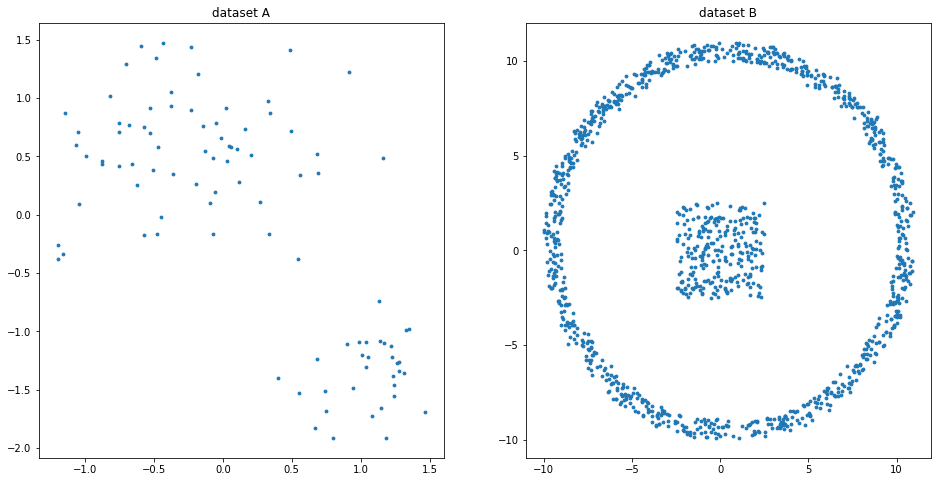
x1, y1, x2, y2 = synthData()
X1 = np.array([x1, y1]).T
X2 = np.array([x2, y2]).T

class kMeans(Distance):
def __init__(self, K=2, iters=16, seed=1):
super(kMeans, self).__init__()
self._K = K
self._iters = iters
self._seed = seed
self._C = None
def _FNC(self, x, c, n):
# for each point,
# find the nearest center
cmp = np.ndarray(n, dtype=int)
for i, p in enumerate(x):
d = self.distance(p, self._C)
cmp[i] = np.argmin(d)
return cmp
def pred(self, X):
# prediction
n, dim = X.shape
np.random.seed(self._seed)
sel = np.random.randint(0, n, self._K)
self._C = X[sel]
cmp = self._FNC(X, self._C, n)
for _ in range(self._iters):
# adjust position of centroids
# to the mean value
for i in range(sel.size):
P = X[cmp == i]
self._C[i] = np.mean(P, axis=0)
cmp = self._FNC(X, self._C, n)
return cmp, self._C
%%time
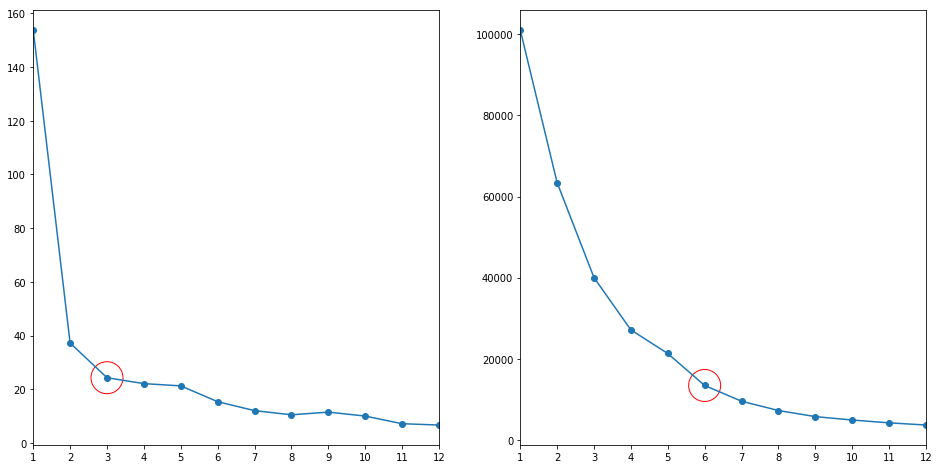
# elbow method
Cs = 12
V1 = np.zeros(Cs)
V2 = np.zeros(Cs)
D = Distance()
for k in range(Cs):
kmeans = kMeans(K=k + 1, seed=6)
fnc1, C1 = kmeans.pred(X1)
fnc2, C2 = kmeans.pred(X2)
for i, [c1, c2] in enumerate(zip(C1, C2)):
d1 = D.distance(c1, X1[fnc1 == i])**2
d2 = D.distance(c2, X2[fnc2 == i])**2
V1[k] += np.sum(d1)
V2[k] += np.sum(d2)
Wall time: 5.35 s
%%time
iters = 20; seed = 6
K1 = 3
kmeans1 = kMeans(K1, iters, seed)
fnc1, C1 = kmeans1.pred(X1)
K2 = 6
kmeans2 = kMeans(K2, iters, seed)
fnc2, C2 = kmeans2.pred(X2)
Wall time: 862 ms
